Identification of levoglucosan degradation pathways in bacteria and sequence similarity network analysis
Por um escritor misterioso
Descrição
Publications - Scott lab
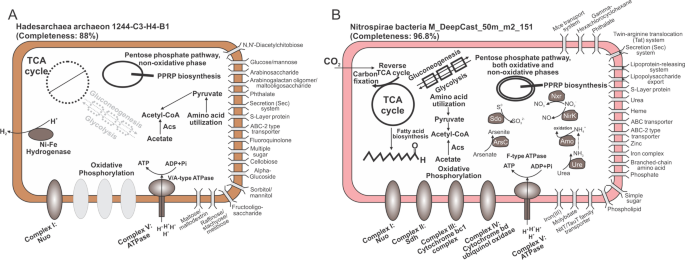
METABOLIC: high-throughput profiling of microbial genomes for functional traits, metabolism, biogeochemistry, and community-scale functional networks, Microbiome

Identification of levoglucosan degradation pathways in bacteria and sequence similarity network analysis

FAD-dependent C-glycoside–metabolizing enzymes in microorganisms: Screening, characterization, and crystal structure analysis

Production of levoglucosan (LG) by pyrolysis of cellulose, and

FAD-dependent C-glycoside–metabolizing enzymes in microorganisms: Screening, characterization, and crystal structure analysis
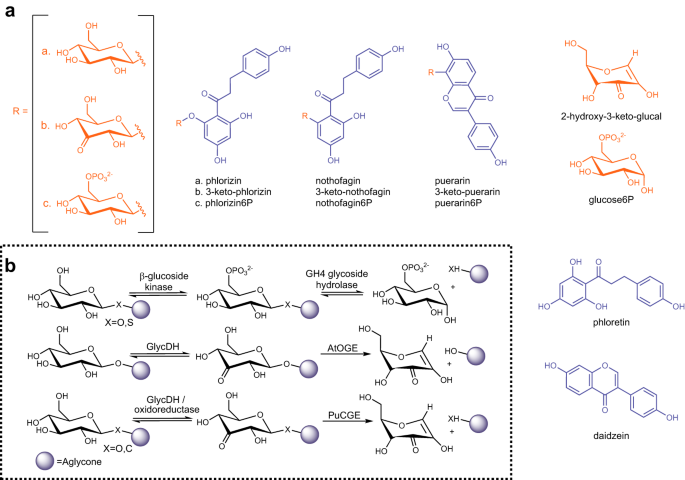
Enzymatic β-elimination in natural product O- and C-glycoside deglycosylation

PDF) ChemInform Abstract: Pathways for Degradation of Lignin in Bacteria and Fungi
Conversion of levoglucosan into glucose by the coordination of four enzymes through oxidation, elimination, hydration, and reduction
Archives of Microbiology

Inhibitor tolerance and bioethanol fermentability of levoglucosan-utilizing Escherichia coli were enhanced by overexpression of stress-responsive gene ycfR: The proteomics-guided metabolic engineering - ScienceDirect
Archives of Microbiology
Prof Spencer Williams

Isolation and Characterization of Levoglucosan-Metabolizing Bacteria
de
por adulto (o preço varia de acordo com o tamanho do grupo)